In Stata 16, you can embed and execute Python code from within Stata. Stata’s new pythoncommand provides a suite of subcommands allowing you to easily call Python from Stata and output Python results within Stata.
You can invoke Python interactively or in do-files and ado-files so that you can leverage Python’s extensive language features. You can also execute a Python script file (.py) directly through Stata.
Additionally, the Stata Function Interface (sfi) Python module is included. It provides a bidirectional connection between Stata and Python. It allows you to interact Python’s capabilities with Stata’s core features. Within the module, classes are defined to provide access to Stata’s current dataset, frames, macros, scalars, matrices, value labels, characteristics, global Mata matrices, and more.
Stata supports both Python 2 and Python 3 starting from Python 2.7. You can choose which one to bind to from within Stata.
Let’s see it work
The first time you call python in Stata, Stata will search for Python installations on the system and choose the one with the highest version. Once Stata finds the candidate with the highest version, it will save that information to use in the future. You can then start your Python journey within Stata. Next we will show you how to invoke Python from Stata.
Invoke Python interactively
You can type python in the Stata Command Window to enter the Python environment. Think of this as an interactive Python shell. You can use it much like you can use Mata (Stata’s built-in matrix programming language) interactively. For example, you could type
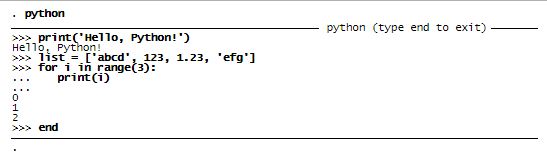
Embed Python code in a do-file
It is easy to embed Python code in a do-file. All you need to do is place the Python code within a python and end block.
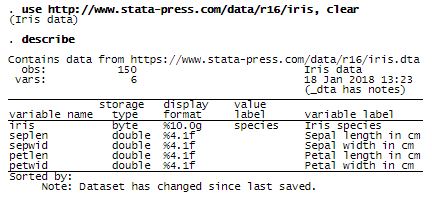
We will use the famous Iris dataset as an illustration. This dataset is used in Fisher’s (1936) article. Fisher obtained the Iris data from Anderson (1935). The data consist of four features measured on 50 samples from each of three Iris species. The four features are the length and width of the sepal and petal. The three species are Iris setosa, Iris versicolor, and Iris virginica.
Our goal is to build a classifier using those features to detect the Iris type. Here we will use the Support Vector Machine (SVM) classifier within thescikit-learn Python package to achieve this goal. Note that you need to install the Matplotlib, sklearn, and NumPy packages in your current Python installation to run the following example. Before using Matplotlib with Stata, you may need to set the backend for different Python installations. See Using Matplotlib with Stata for a discussion about calling Matplotlib from Stata. We put the following code in the Do-file Editor and execute it:
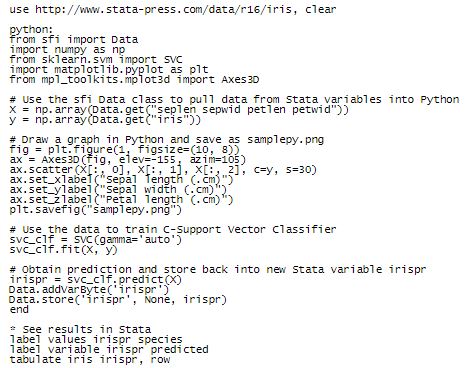
In the code above, we did the following:
Imported all the modules, functions, and objects we were going to use.
Loaded the four features and the species type into two NumPy arrays X and y, respectively. Note that for simplicity, we do not split our dataset here and we used all the instances as our training samples.
Drew a 3D scatter plot using Matplotlib.
Built an SVC classifier for classification defined in the sklearn package and fit the model.
Made predictions for the dataset using the trained classifier.
Stored the predictions into a new variable irispr in Stata.
Back in Stata, attached the value label of iris onto irispr, and used the tabulate command to display a classification table.
We saved the above code in samplepy.do and ran
. do samplepy
which produced the following image and output:
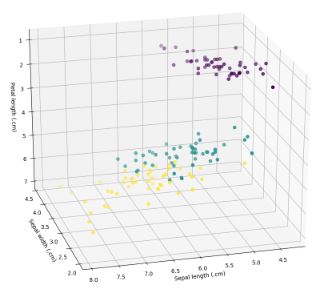
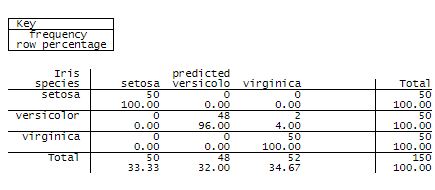
The above table shows that 2 Iris versicolor observations were misclassfied as Iris virginica, and no Iris setosa or Iris virginica were misclassified.
Embed Python code in an ado-file
Python code can be embedded and executed in ado-files too. Below, we create a new command mysvm in mysvm.ado to illustrate this purpose. mysvmexpects a label variable to be specified first followed by a list of feature variables along with a predict() option specifying the name of the variable where the prediction will be stored.

In the above ado-file, we defined the classifier within the Python function dosvm(), which took the species type variable, the four feature variables, and the new variable storing the predictions as arguments. We called the Python function using the python: istmt syntax in the ado-code.
To produce the same output as above, we can type


